Charting New Waters - MGnify Genomes Marine Catalogue v2.0 Released
We are excited to announce the release of an updated version of the MGnify Genomes marine catalogue.
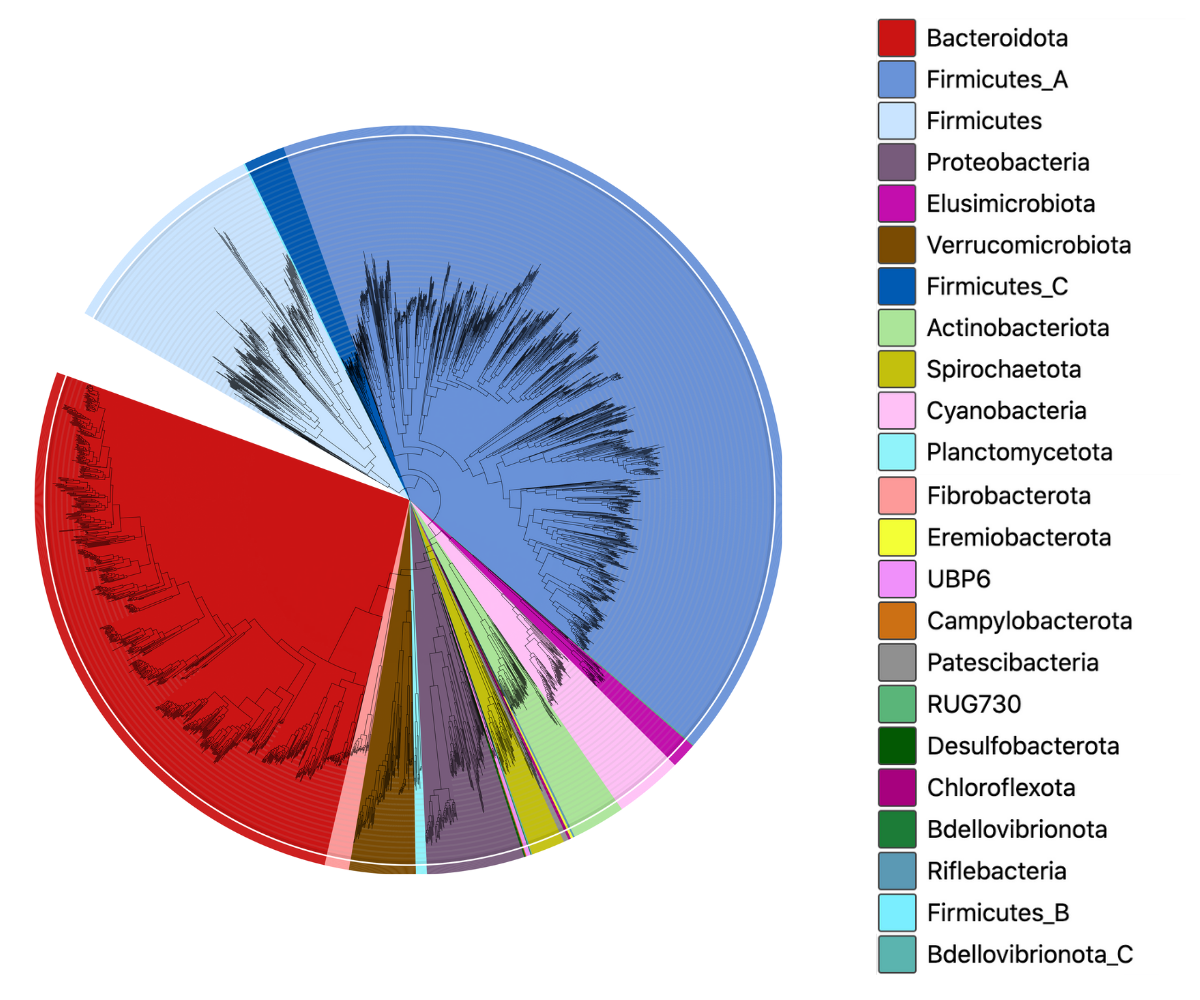
This latest release (v2.0) of the marine catalogue contains data from 1628 studies, including genomes from major sampling expeditions such as Tara Oceans, Malaspina, GO-Ship, and Geotraces, amongst others.

 MGnify is delighted to announce the release of our latest
MGnify is delighted to announce the release of our latest 


 The microbial population (or microbiome) of the human gut is involved in a wide range of important processes, such as digestion, production of vitamins and other nutrients, detoxification, protection from pathogens, and helping to shape the host immune system. Gut microbial communities represent substantial reservoirs of genetic and metabolic diversity: different people have different types of microorganisms in their gut, and community composition can change over time or with diet.
The microbial population (or microbiome) of the human gut is involved in a wide range of important processes, such as digestion, production of vitamins and other nutrients, detoxification, protection from pathogens, and helping to shape the host immune system. Gut microbial communities represent substantial reservoirs of genetic and metabolic diversity: different people have different types of microorganisms in their gut, and community composition can change over time or with diet.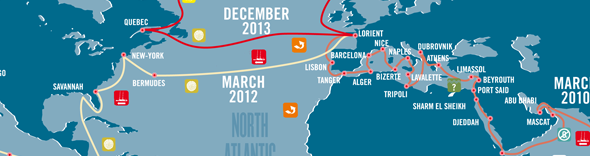 Plankton ecosystems contain a phenomenal reservoir of life: more than 10 billion organisms inhabit every litre of oceanic water, including viruses, prokaryotes, unicellular eukaryotes (protists), and metazoans.
Plankton’s importance for the earth’s climate is at least equivalent to that of the rainforest. Yet only a small fraction of organisms that compose it have been classified and analysed.
Plankton ecosystems contain a phenomenal reservoir of life: more than 10 billion organisms inhabit every litre of oceanic water, including viruses, prokaryotes, unicellular eukaryotes (protists), and metazoans.
Plankton’s importance for the earth’s climate is at least equivalent to that of the rainforest. Yet only a small fraction of organisms that compose it have been classified and analysed.